I am transitioning, to my new website: jeffareslab.org
♠ How is genome evolution influenced by genome and cell function? I use a variety of data and computational techniques to describe how genomes change but still continue to produce a working molecular machinery for the cell. I apply methods from molecular evolution, population genomics, comparative genomics, functional genomics (also tinkering and thinking).
-

- Each line and circle show the estimated heritability of a trait. The black bar shows how much CNVs contribute. The plot at the right shows how much we would attribute to CNVs only because of linkage to causal SNPs (not much at all).
♠ Current research
I am exploring the genomic and phenotypic diversity of fission yeast, both to understand this species, and to describe how genome evolution is influenced by genome and cell function (read about this).
♠ Previous topics
» origin and early evolution of life (read about this)
» evolution of introns (read more)
» genomic diversity of malaria parasites (Plasmodium) (read more)
» plant developmental biology (read more)
♠ Genomic and phenotypic diversity of the fission yeast
Part one: The fission yeast Schizosaccharomyces pombe is a principle model for cell a biology. Its an excellent model for population genomics, because genomic factors can be linked to the cell biology that they influence. I recently published an article in Nature Genetics that described the genomic diversity of all known wild strains, described their evolutionary history and characterised 70 quantitative traits, allowing us to conduct the first 220 genome-wide association studies (GWAS) for this species.
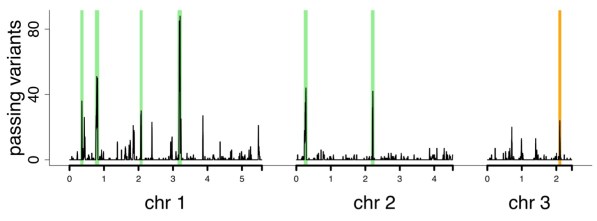
Part two: We then looked at structural variation (large duplications/deletions, inversions and translations). We showed that not only do structural  variants contribute considerably to heritable traits, they are transient within populations (they change rapidly). We were surprised, but the evidence is incontrovertible. This will be out in Nature Communications soon, but you can read the pre-print on bioRxiv.
variants contribute considerably to heritable traits, they are transient within populations (they change rapidly). We were surprised, but the evidence is incontrovertible. This will be out in Nature Communications soon, but you can read the pre-print on bioRxiv.
Pombe makes wine: I have since shown that some of these wild strains make good wine (with Santiago Benito Saez, a scientist and a winemaker), and that structural variants contribute strongly to quantitative traits and to intrinsic reproductive isolation (now on bioRxiv). I am also exploring how the Hermes transposon can be used to describe the functional elements in the genome.
 ♠ The genomic diversity of malaria parasites
♠ The genomic diversity of malaria parasites
While at the Wellcome Trust Sanger Institute I described one of the first genome-wide diversity data sets for the malaria parasite Plasmodium falciparum. With two other papers, we were featured on the cover of Nature Genetics.
♠ The origin and early evolution of life
What can we know about events that occurred more than a billion years ago? With David Penny and Anthony Poole, we studied the complexity of modern RNA use to ‘retrodict’ the final complexity of the RNA world, describing the hypothetical species Riborgis eigensis. Commentaries about our research on this topic appeared here:
Evolution: Genome data shake tree of life. Science 1998 May;80:672.
Making Life Simple. New Scientist 1999 Jan;2169:34-7, and also here.
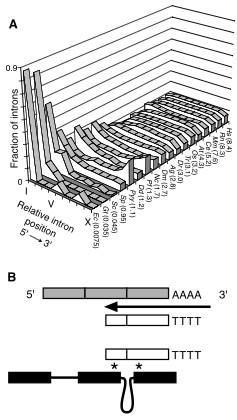 ♠ Intron evolution With David Penny, Tobias Mourier and others, I joined in the debate about intron gain, and intron loss in eukaryote genomes. Finding signals of intron loss in eukaryote genomes (Science 2003), that intron loss is influenced by a variety of facets of molecular biology (Trends in Genetics 2006) and that rapidly regulated genes are intron poor (Trends in Genetics 2008).
♠ Intron evolution With David Penny, Tobias Mourier and others, I joined in the debate about intron gain, and intron loss in eukaryote genomes. Finding signals of intron loss in eukaryote genomes (Science 2003), that intron loss is influenced by a variety of facets of molecular biology (Trends in Genetics 2006) and that rapidly regulated genes are intron poor (Trends in Genetics 2008).
♠ Plant developmental biology
During my PhD with Bruce Veit, I studied mei2-like genes, the Arabidopsis genome, and the terminal-ear1 gene in maize.